Sex Chromosome Evolution Simulator
Explore how homologous autosomes diverge into distinct sex chromosomes through four evolutionary stages. Click on each stage to see the progression.
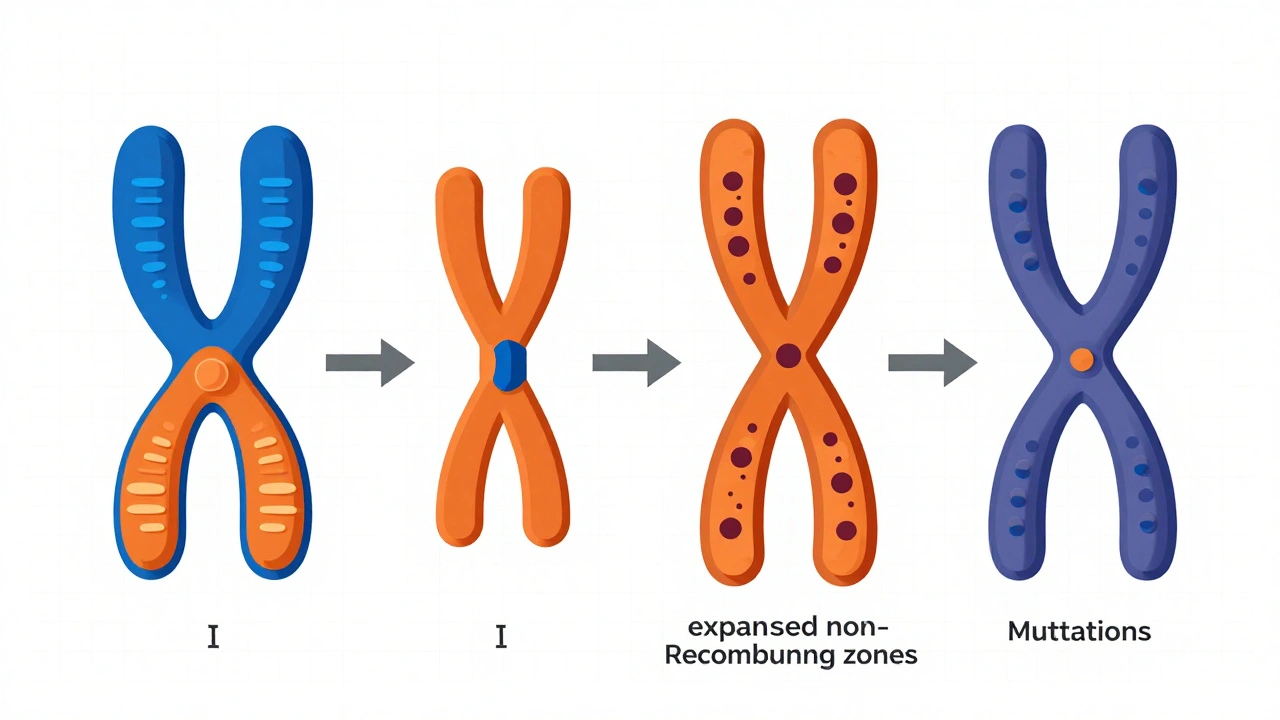
Origin
Identical autosomes with a new sex-determining gene appearing on one chromosome.
First Inversion
Recombination stops around the sex-determining gene due to chromosomal inversion.
Expansion
More inversions expand the non-recombining region. Deleterious mutations accumulate.
Degeneration
The Y chromosome degenerates significantly, losing most genes while X remains stable.
Visual Simulation
Recombination Status:
Active recombination between homologous regionsGene Loss (Y Chromosome):
No significant gene loss yetStage I: Origin
An ordinary pair of autosomes acquires a sex-determining locus. The chromosomes are still mostly identical and recombine freely during meiosis. This is the starting point for sex chromosome evolution.
For a long time, scientists thought sex chromosomes were just old autosomes that got stuck in a bad relationship. They believed the process was simple: two identical chromosomes pick a sex-determining gene, stop swapping DNA, and one slowly rots away while the other stays healthy. It’s a clean, linear story. But if you look at the actual data from fungi to humans, that story falls apart. The reality is messier, faster, and far more dynamic.
Semantic analysis of Sex Chromosome Evolution, the process by which homologous autosomes diverge into distinct sex-determining structures through recombination suppression and mutation accumulation reveals that these genetic elements are not static endpoints but active participants in genomic conflict. We need to understand how this divergence happens, why it varies so wildly across species, and what recent discoveries tell us about the forces driving these changes.
The Classical Model: Four Stages of Divergence
To understand where we are today, we have to look at the baseline theory. In 1999, researchers Bruce Lahn and David Page published a landmark study in Science that mapped out the history of human X and Y chromosomes. Their model suggested that about 300 million years ago, an ordinary pair of autosomes acquired a sex-determining locus. This triggered a four-stage process of divergence.
- Stage I: A sex-determining gene appears on one chromosome. The chromosomes are still mostly identical and recombine freely.
- Stage II: Recombination stops around the sex-determining gene due to chromosomal inversions. This creates a small non-recombining region.
- Stage III: More inversions occur, expanding the non-recombining region. Deleterious mutations begin to accumulate on the proto-Y because they can't be purged by recombination.
- Stage IV: The Y chromosome degenerates significantly, losing most of its genes, while the X remains stable because females (XX) can still recombine their X chromosomes.
This model works well for mammals. It explains why the human Y chromosome is tiny compared to the X. However, treating this as a universal rule ignores the vast diversity seen in nature. Not all lineages follow this slow, steady decay.
The Role of Recombination Suppression
The engine of sex chromosome differentiation is Recombination Suppression, the prevention of genetic exchange between homologous chromosomes during meiosis, often caused by structural inversions. When two chromosomes swap segments during meiosis, they repair errors and shuffle beneficial traits. If they stop swapping, they drift apart genetically.
Inversions-where a segment of DNA flips orientation-are the primary mechanism for stopping this swap. If a region on the Y chromosome is inverted relative to the X, the chromosomes cannot align properly during meiosis. Without alignment, no crossing over occurs. Each new inversion adds another block of DNA to the "no-trade zone." Over time, the Y chromosome becomes a genetic island, isolated from the repairing power of the X.
| Region Type | Recombination Status | Evolutionary Age |
|---|---|---|
| Pseudoautosomal Region (PAR) | Active | Ongoing |
| Differentiation Zone 1 | Suppressed | Ancestor of H. lupulus |
| Differentiation Zone 2 | Suppressed | Modern H. lupulus populations |
The genus Humulus, a plant genus including hops, used as a model for studying young sex chromosome systems with distinct recombination zones provides a perfect snapshot of this process. Researchers identified three distinct regions on Hop sex chromosomes. One region still recombines (the pseudoautosomal region), while two others stopped recombining at different times in the past. This proves that differentiation is stepwise, not instantaneous.
Exceptions to the Rule: Ephemerality and Turnover
If the classical model were absolute, every lineage would eventually end up with a tiny, degenerate Y or W chromosome. Instead, we see something surprising: Sex Chromosome Turnover, the replacement of one sex-determination system with another, often involving new chromosomes or genes. Sex chromosomes are often ephemeral. They don’t last forever.
Turnover happens when a new mutation arises that takes over sex determination. This can happen on a completely different autosome. Suddenly, the old X and Y become ordinary autosomes again, and a new pair starts the divergence process. This is common in fish, amphibians, and reptiles. For example, some species of fish switch between XY and ZW systems within closely related groups. This plasticity suggests that sex determination is not a fixed trait but a labile one, subject to rapid evolutionary change.

Mutation vs. Selection: What Drives Degeneration?
Why does the Y chromosome degenerate? The traditional view is sexual selection. Genes beneficial to males (like those for antlers or manes) hitchhike onto the Y chromosome near the sex-determining gene. Over time, the Y accumulates male-specific genes. But there’s a competing hypothesis: degeneration might be driven simply by the inability to purge deleterious mutations.
This idea has been tested in fungi. Fungi offer a unique comparison because some are diploid (two copies of each chromosome) for most of their life, while others are haploid (one copy). If sexual selection drives differentiation, both should show differentiated mating-type chromosomes. If mutation-purging capacity drives it, only diploids should show differentiation because haploids cannot hide harmful mutations-they are exposed immediately to natural selection.
Empirical evidence supports the latter. Diploid fungi exhibit differentiated mating-type chromosomes, similar to heteromorphic sex chromosomes in animals. Haploid fungi do not. This suggests that genome structure and the ability to shelter mutations play a bigger role in early differentiation than previously thought. Sexual selection may amplify differences later, but the initial split might be largely mechanical.
Hemizygosity and Dosage Compensation
Once sex chromosomes differentiate, a new problem arises: dosage imbalance. In XY systems, males have one X and one Y, while females have two Xs. If X-linked genes are expressed twice as much in females, this could be lethal. Nature solves this with Dosage Compensation, mechanisms that equalize gene expression levels between sexes despite differing numbers of sex chromosomes.
In mammals, one X chromosome in females is randomly silenced and condensed into Barr body. This ensures that both males and females express X-linked genes at roughly the same level. In birds (ZW system), the mechanism is different; males (ZZ) downregulate one Z chromosome slightly, rather than silencing it completely. These mechanisms are critical for survival and evolve quickly after sex chromosomes diverge.
Another consequence of hemizygosity is Meiotic Silencing, the transcriptional silencing of unsynapsed chromatin regions during meiosis to prevent inappropriate recombination. During male meiosis, the parts of the X and Y that don’t match up are silenced. This prevents the cell from detecting them as damaged DNA and triggering apoptosis. This specialized chromatin state affects gene expression and evolution rates on sex chromosomes.

Impact on Speciation and Hybrid Incompatibility
Sex chromosomes are hotspots for speciation. Haldane's Rule, the principle that if one sex of hybrid offspring is absent, rare, or sterile, it is usually the heterogametic sex (XY or ZW) states that hybrids of the heterogametic sex (males in mammals, females in birds) are more likely to be sterile or inviable than the homogametic sex.
This happens because recessive alleles on the X or Z chromosome are exposed in the heterogametic sex. In autosomes, harmful recessive alleles can be hidden by a dominant healthy allele. On the single X in males, there is no backup. If the X chromosome from Species A doesn’t work well with the autosomes from Species B, the male hybrid fails. This makes sex chromosomes major contributors to reproductive isolation between species.
Convergent Patterns Across Systems
Despite the diversity, XY and ZW systems show convergent evolution. Both start as homologous autosomes. Both suppress recombination via inversions. Both undergo degeneration of the heterogametic chromosome (Y or W). Both develop dosage compensation. The specific genes involved differ, but the trajectory is remarkably similar.
This convergence suggests strong selective pressures shaping sex chromosome architecture. Whether it’s mammals with XY systems or butterflies with ZW systems, the underlying logic of genetic isolation and mutation accumulation holds true. However, the rate of change varies. Some systems are ancient and stable (mammals), while others are young and fluid (fish, plants).
Future Directions in Research
As of 2026, research focuses on understanding the triggers for turnover and the role of epigenetic factors in dosage compensation. New genomic tools allow scientists to track inversion events in real-time in rapidly evolving species. Understanding these dynamics helps us grasp not just sex determination, but broader principles of genome evolution, including how genomes balance stability with adaptability.
The story of sex chromosomes is no longer just about decay. It’s about dynamic interplay between mutation, selection, and structural change. By looking beyond the classical model, we gain a richer understanding of how life diversifies at the most fundamental level.
Do all species have XY or ZW sex chromosomes?
No. Many species use environmental sex determination (temperature-dependent in reptiles), haplodiploidy (bees and ants), or other genetic systems. Even among genetic systems, the chromosomes involved vary widely. Some fish and amphibians have multiple pairs of sex chromosomes.
Why doesn't the Y chromosome disappear entirely?
The Y chromosome retains essential genes for male fertility and development. While it has lost many ancestral genes, it has evolved testis-specific amplifications and unique regulatory mechanisms to maintain function. Complete loss would require a total shift in sex determination mechanism.
What causes recombination suppression?
Chromosomal inversions are the primary cause. When a segment of DNA flips orientation on one chromosome, it cannot align properly with the other during meiosis, preventing crossover. Other mechanisms include sequence divergence and epigenetic modifications.
Is sex chromosome evolution always gradual?
Not necessarily. While the classical model suggests gradual expansion of non-recombining regions, recent studies show punctuated equilibrium patterns. Large inversion events can cause rapid shifts. Additionally, sex chromosome turnover can reset the clock entirely.
How do fungi help us understand sex chromosome evolution?
Fungi provide a natural experiment comparing haploid and diploid life cycles. Diploid fungi show differentiated mating-type chromosomes, supporting the hypothesis that mutation-purging capacity drives early differentiation. Haploid fungi do not, highlighting the role of genome structure.